Sie befinden sich hier
Inhalt
mRNA expression dynamics in innate immune cells
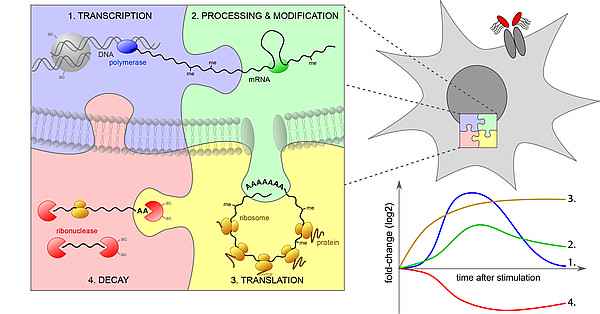
When innate immune cells react to inflammatory stimuli, gene expression is regulated rapidly at multiple levels: Transcription, mRNA processing and export, translation and mRNA decay all contribute to the complex dynamics of the response. Often, different levels of mRNA expression are tightly connected: Many cytoplasmic RNA-binding proteins, for example, affect both mRNA stability and translation. Likewise, mRNA modifications that are deposited co-transcriptionally in the nucleus regulate the fate of an mRNA in the cytoplasm. As mRNA expression is far from steady state conditions during the inflammatory response, different expression parameters additionally interact due to logistic constraints along the way from transcription to decay. We develop both experimental and bioinformatics/statistical methods to disentangle the contribution of individual steps of gene expression and identify key post-transcriptional regulators of the inflammatory response.
Selected publications
Schott, J., Reitter, S., Lindner, D., Grosser J., Bruer M., Shenoy, A., Geiger T., Mathes A., Dobreva G., Stoecklin G. Nascent Ribo-Seq measures ribosomal loading time and reveals kinetic impact on ribosome density. Nat Methods 18, 1068–1074 (2021).
Haneke K, Schott J, Lindner D, Kruse Hollensen A, Kroun Damgaard C, Mongis C, Knop M, Palm W, Ruggieri A, Stoecklin G. CDK1 couples proliferation with protein synthesis. J Cell Biol. 2020 Mar 2;219(3):e201906147.
Schott J, Reitter S, Philipp J, Haneke K, Schäfer H, Stoecklin G. Translational regulation of specific mRNAs controls feedback inhibition and survival during macrophage activation. PLoS Genet. 2014;10(6):e1004368.
Leppek K, Schott J, Reitter S, Poetz F, Hammond MC, Stoecklin G. Roquin promotes constitutive mRNA decay via a conserved class of stem-loop recognition motifs. Cell 2013;153:869-81.
Schott J, Stoecklin G. Networks controlling mRNA decay in the immune system. Wiley Interdiscip Rev RNA 2010;1:432-56.
Kontextspalte
Project Group Leader