Sie befinden sich hier
Inhalt
Transcriptional and epigenetic control of cardiovascular development and disease
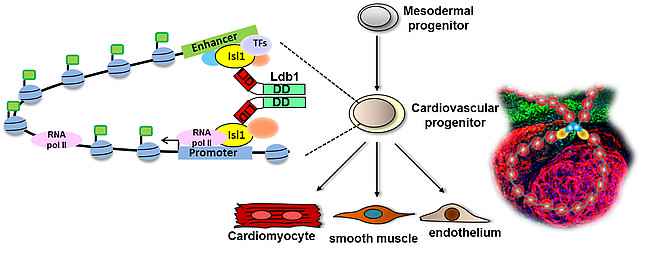
Heart formation is a complex morphogenetic process, involving specification and differentiation of cardiovascular progenitor cells into distinct cell lineages, such as cardiomyocytes, smooth muscle cells and endothelial cells. All these cell types contribute to the structural, mechanical and electrical properties of the heart. On a molecular level, a large set of transcription factors, chromatin modifiers and signalling molecules are integrated in regulatory networks that drive the establishment of stable gene expression patterns characteristic for these distinct cardiac cell types, while also allowing for a certain level of plasticity, for example to enable restoration of tissue homeostasis following injury and other environmental challenges.
In the last years we have identified key factors involved in cardiac lineage specification, progenitor cell expansion, cell fate decisions and differentiation. We uncovered Ajuba as an important downstream effector of retinoic acid signaling that regulates the cardiac progenitor pool (Witzel et al., 2012). Additionally, we found that a complex between the adaptor molecule Ldb1 and the key cardiac transcription factor Isl1 plays a key role in orchestrating a network for heart-specific transcriptional regulation and coordination in three-dimensional space during cardiogenesis (Caputo et al., 2015).
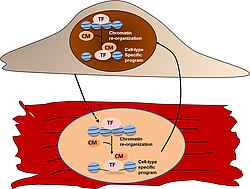
Moreover, using genome-wide approaches we uncovered epigenetic modifiers that are crucial for chromatin reorganization and transcriptional programming resulting in cardiac lineage commitment. We are currently studying their role in genetic reprogramming in cardiovascular development, homeostasis and maladaptive remodelling in cardiovascular diseases. A major focus lies on understanding the crosstalk between cellular metabolism and alterations in the epigenetic landscape during these processes.
Pathways and players in zebrafish heart development and regeneration
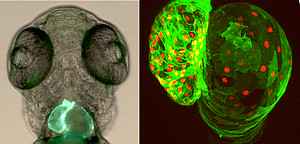
The zebrafish has emerged as a powerful model system to study heart development and disease, due to the large body of evidence for genetic conservation between zebrafish and mammalian cardiogenesis. Zebrafish combines several advantages, such as external development and transparency of the embryo, which allow live imaging of the complex morphogenetic events occurring during heart development. In addition, the zebrafish exhibits an ability to regenerate its heart, and thus, can provide critical insights into how regeneration is achieved or might be stimulated. We are using CRISPR/Cas9-mediated genome editing technology to study the function of selected genes in cardiac morphogenesis and function and to generate cardiovascular disease models for gene variants identified in high-throughput sequencing projects of patients with congenital heart disease (CHD). Furthermore we analyse transcriptomic and epigenomic alterations during zebrafish development and regeneration with the aim of identifying key pathways and players in cardiac regeneration.
Endothelial-cardiomyocyte interactions in cardiovascular development and disease
Endothelial-cardiomyocyte communications enable these cell types to coordinate their activity and are essential for cardiac morphogenesis, postnatal cardiac growth, contractile performance and rhythmicity. Endothelial cells supply cardiomyocytes not only with oxygenated blood but also secrete factors that control cardiomyocyte organization, proliferation, survival and contraction. While endothelial cells instruct cardiomyocyte behaviour, cardiomyocytes conversely secrete factors that control endothelial cell function. We study the molecular mechanisms leading to transcriptional and epigenetic programming elicited by inductive signals as a result of endothelial-cardiomyocyte interactions.
Epigenetics in cancer initiation and progression
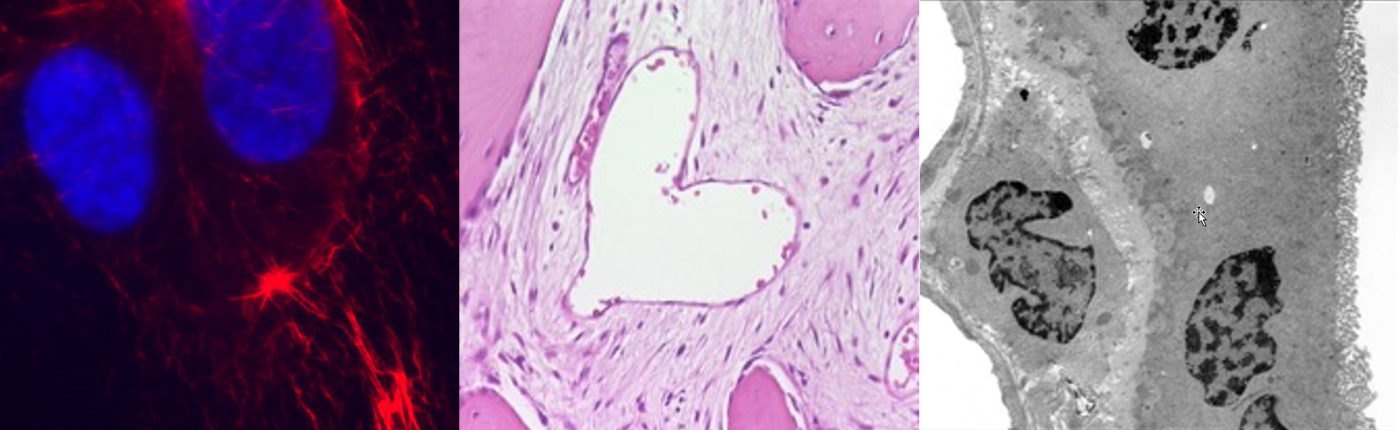
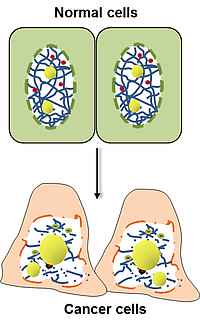
Changes of nuclear morphology are a diagnostic hallmark of tumors, but the mechanisms that mediate them are poorly understood. We found several components of the nuclear envelope and the nucleoskeleton to act as tumor suppressors, thus providing a molecular link between aberrant nuclear architecture, and the malignant phenotype. Genetic loss of these components leads to spontaneous tumor formation with high incidence. We are currently studying their role in higher-order chromatin organization, maintenance of genome stability and transcriptional regulation. Additionally, we analyze the influence of nuclear structure alterations on cytoskeletal organization, cell migration and invasion.